The awesome power of yeast evolutionary genetics: New genome sequences and strain resources for the Saccharomyces sensu stricto genus
Devin R. Scannell*, Oliver A. Zill*, Antonis Rokas, Celia Payen, Maitreya J. Dunham, Michael B. Eisen, Jasper Rine, Mark Johnston, Chris Todd Hittinger
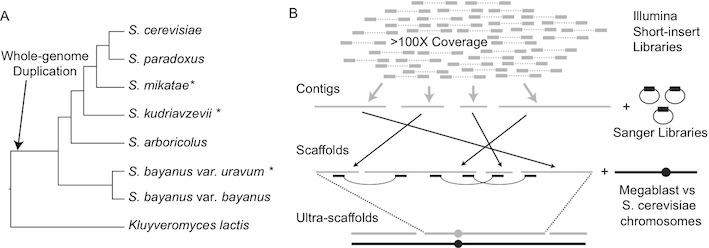
* Co-first authors listed alphabeticallyHigh-quality, well-annotated genome sequences and standardized laboratory strains fuel experimental and evolutionary research. We present improved genome sequences of three species of Saccharomyces sensu stricto yeasts: S. bayanus var. uvarum (CBS 7001), S. kudriavzevii (IFO 1802 T), S. kudriavzevii (ZP 591), and S. mikatae (IFO 1815 T), and describe their comparison to the genomes of S. cerevisiae and S. paradoxusThe new sequences, derived by assembling millions of short DNA sequence reads together with previously published Sanger shotgun reads, have vastly greater long-range continuity and far fewer gaps than the currently available genome sequences, and enable previously impossible analyses. New gene predictions defined a set of 5,261 protein-coding orthologs across the five most commonly studied Saccharomyces yeasts, enabling a re- examination of the tempo and mode of yeast gene evolution and improved inferences of species-specific gains and losses. To facilitate experimental investigations, we generated genetically marked, stable haploid strains for all three of these Saccharomyces species. These nearly complete genome sequences and the collection of genetically marked strains provide a valuable toolset for comparative studies of gene function, metabolism, and evolution, and render Saccharomyces sensu stricto the most experimentally tractable model genus.
Paper now available (Open Access) in the inaugural issue of G3!